Junk DNA: A Journey Through the Dark Matter of the Genome (7 page)
Read Junk DNA: A Journey Through the Dark Matter of the Genome Online
Authors: Nessa Carey

There’s a possible scenario that has worried some scientists. Anyone who receives a pig heart will inevitably be receiving immunosuppressive drugs to prevent rejection of the foreign organ. Reactivation of endogenous retroviruses is more likely when individuals are immunosuppressed. Human systems have evolved in part to control the endogenous retroviruses that have been in our genome since we evolved. But they may not be as efficient at controlling the ones hiding in the pig genome. This theoretically could mean that the endogenous retroviruses could escape from the pig heart and attack and enter other cells in the human recipient. From there, they might even escape into the wider population.
More recent data have suggested that the risk of this happening has perhaps been overstated in the past,
11
but it’s certainly an area of junk DNA that will require close scrutiny if xenotransplantation is to become a reality.
Other repeated sequences in the genome can cause health problems more directly. There are some parts of the genome where large sections, sometimes hundreds of thousands of base pairs in length, were duplicated relatively recently during human evolution. The ‘original’ and the ‘duplicate’ may end up in very different parts of the genome, even on different chromosomes from one another.
These regions can cause problems when eggs or sperm are being formed. During this formation, there is a very important stage where chromosomes undergo a process called crossing-over. A chromosome inherited from your mother pairs up with the equivalent chromosome inherited from your father, and they swap bits of DNA between the two. It’s a way of increasing the amount of variation in the gene pool, by mixing up combinations of genes. If there are two parts of the genome that look very similar because
of repeat sequences but which are not actually a matching pair of chromosomes, this crossing-over may occur between regions of the genome that aren’t meant to swap material. The consequence may be that eggs or sperm are produced that have extra sections of DNA, or are missing critical regions.
12
This can lead to disease in individuals who inherit these genomic defects. One example is Charcot-Marie-Tooth disease, where there are defects in the nerves that transmit sensation and control motor functions.
13
Another is Williams-Beuren syndrome, a condition characterised by developmental delay, relative shortness, a range of unusual behavioural traits combined with mild learning disability, and long-sightedness.
14
The duplicated regions in the genome that give rise to the problems during crossing-over often contain multiple protein-coding genes. It’s probably not surprising that the symptoms in patients affected by abnormal crossing-over are often quite complex. It’s likely that more than one pathway is affected by the change in the number of multiple genes.
It might seem odd that these duplicated regions have been retained during human evolution, if they can give rise to such problems. But in reality, most of the time the cells that form eggs and sperm perform crossing-over really well, and don’t mix up the wrong parts of chromosomes. The duplications have also acted as a way that the human genome has been able to increase the numbers of certain genes quite rapidly, in evolutionary terms. This can be useful. The ‘spare’ copy may act as the raw material for evolutionary adaptation. A few changes to the protein-coding gene sequence can create a protein with a related but discrete function from the original. This may be how the large family of genes that allows mammals to detect a huge range of different smells evolved.
15
It’s another example of the parsimony with which the human genome has evolved, adapting existing genes and proteins, rather than starting from scratch. A genomic two-for-one offer.
From guilt to innocence via junk DNA
Most of the junk repetitive DNA that we have considered so far in this chapter is formed of quite large units. These tend to be at least 100 base pairs in length and are frequently much longer. That’s partly why they account for so much of the genome. But there are other junk repetitive units that are much smaller, based on repeats of just a few base pairs. These are called simple sequence repeats. We already met a few examples of these in the exploration of Fragile X syndrome, Friedreich’s ataxia and myotonic dystrophy. In each of these cases, three-base-pair sequences were repeated a number of times, and reached their maximum in patients with the disorders.
Repeats of short motifs account for about 3 per cent of the human genome. They are very variable between individuals. Let’s consider an arbitrary repeat of two base pairs, say GT, at a particular position on chromosome 6. I may have inherited eight copies (sequence would be GTGTGTGTGTGTGTGT) on chromosome 6 from my mother and seven copies on chromosome 6 from my father. You, on the other hand, may have inherited ten copies from your mother and four from your father.
These simple sequence repeats have proved to have great usefulness because they are found all over the genome, vary a lot between individuals at each position where they occur in the genome and are easy to detect using cheap, sensitive methods.
Because of these characteristics, such repeats are now used for DNA fingerprinting. This is the process by which blood or tissue samples can be unequivocally associated with a specific individual. This has facilitated paternity testing and revolutionised forensic science. Its applications in the latter have included identification of victims of massacres, convictions of the guilty and exonerations of the innocent, including cases where the wrong person has been in jail for decades. Over 300 people in the United States have been freed after DNA testing established their innocence, nearly 20 of
whom had been on death row at some point during their incarceration.
16
Additionally, in about half of these cases, DNA evidence was able to determine the real guilty party.
Not bad for a bit of junk.
Footnote
a
The four classes are known as SINES (short interspersed repetitive repeats); LINEs (long interspersed elements); LTRs (elements with long terminal repeats); DNA transposons.
5. Everything Shrinks When We Get Old
The movie
Trading Places
, starring Dan Aykroyd, Eddie Murphy and Jamie Lee Curtis, was a huge hit in 1983, grossing over $90 million at the US box office.
1
It’s a convoluted comedy but the premise behind it is the exploration of genes versus environment. Is a successful man successful because of intrinsic merit or because of the environment in which he is placed? The movie comes out firmly on the side of the latter.
A similar phenomenon can happen in our genomes. An individual gene may perform a relatively innocuous role, helping a cell keep on keeping on, so to speak. The gene produces protein at just the right rate to do this job. A major factor in controlling the amount of protein that is produced is the position of the gene on the chromosome.
Now let’s imagine that the gene is transported to a new neighbourhood, like Dan Aykroyd’s character ending up in the slums or Eddie Murphy’s character finding himself transported to a mansion. In this neighbourhood, our transported gene is surrounded by new genomic information, which instructs it to make much higher amounts of protein. The high levels of the protein whip the cell forwards, pushing it to grow and divide much faster than usual. This can be one of the steps that leads to cancer. There’s nothing bad about the gene itself, it’s just in the wrong place at the wrong time.
This process is caused when two chromosomes break in a cell at the same time. When a chromosome breaks, a repair machinery
immediately targets the break and joins the two bits up again. This is usually a pretty slick process. But if two (or more) chromosomes break at the same time, there can be problems. The ends of the chromosomes may become joined up incorrectly, as shown in Figure 5.1. This is how a ‘good’ gene may end up in a ‘bad’ neighbourhood, and begin causing problems. This is particularly an issue because the rearranged chromosomes will be passed on to all daughter cells every time cell division takes place. Probably the most famous example of this mechanism is in a human blood cancer called Burkitt’s lymphoma, where there is a rearrangement between chromosomes 8 and 14. This results in very strong over-expression of a gene
a
that encourages cells to proliferate aggressively.
2
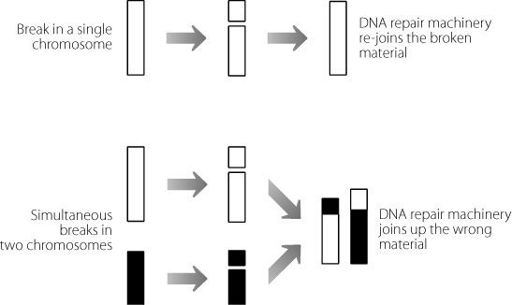
Figure 5.1
In the upper panel a single chromosome breaks and is repaired by the cell. In the lower panel two chromosomes break simultaneously. The cell machinery may be unable to work out which break occurred on which chromosome. The chromosomes may be joined together inappropriately, creating hybrid structures.
Luckily, it’s probably quite rare that two chromosomes break at exactly the same time. More frequently there will be a time difference. So, the machinery that repairs DNA has evolved to act really quickly. After all, the faster it repairs a break, the lower the chance that there will be multiple breaks present at the same time in an individual cell. The DNA repair machinery starts to operate as soon as the cell detects that there is a broken piece of DNA. It does this by having mechanisms to detect the end of the break.
But this creates a whole new set of problems. Our cells contain 46 chromosomes, each of which is linear. In other words, our cells always have 92 chromosome ends, one at each end of a chromosome. The DNA damage machinery has to have a way of distinguishing the perfectly normal ends of chromosomes from the abnormal ends caused by breakages.
DNA shoelaces
The way that cells have solved this is to have special structures on the normal ends of the chromosomes. Are you wearing shoes with laces? If so, have a quick look at those laces. At either end there is a little cap made from metal or plastic. This is called the aglet, and it stops the lace from unravelling and fraying. Our chromosomes have their own aglets, and these are extremely important for maintaining the integrity of our genome.
These chromosomal aglets are called telomeres and they are made from a form of junk DNA that we have known about for many years, plus complexes of various proteins. The telomeric DNA is formed from repeats of the same six base pairs, TTAGGG, repeated over and over again.
3
These stretch for an average of about 10,000 base pairs in total on each end of every chromosome in the umbilical cord blood of a newborn human baby.
4
The telomeric DNA is bound by complexes of proteins that help to maintain the structural integrity.
b
The term telomere really refers to the combination of the junk DNA and its associated proteins. A graphic demonstration of the importance of these proteins was shown by some researchers working in mice in 2007. They knocked out expression of one of the proteins by completely inactivating its gene, and found that the resulting mice embryos died early in development.
c
When the researchers examined the chromosomes in these genetically modified mice, they found that many of them had joined up. The ends had linked up with each other. This was because the DNA repair machinery no longer recognised the telomeres as telomeres. Instead, it reacted as if faced by a whole slew of broken chromosomes and did what it does best. It stuck them together. Unfortunately, by doing so, gene expression became completely disordered. Eventually the chromosomes and cells became so dysfunctional that they triggered a type of cellular suicide,
d
halting development completely.
There is also another feature of the telomeres that is of major interest in biology and human health. Back in the 1960s, researchers were studying how cells divide in the laboratory. They didn’t work with cancer cell lines, as these are derived from cells that have become immortal through abnormal changes. Instead, they studied a kind of cell known as a fibroblast. Fibroblasts are found in a wide range of human tissues. They secrete something called the extracellular matrix, a sort of thick wallpaper paste that holds
the cells in position. It’s relatively easy to take a biopsy, for example from skin, and isolate the fibroblasts. These will grow and divide in culture. What the researchers discovered all those years ago was that the cells wouldn’t keep dividing forever. There came a point when they stopped dividing, even when supplied with all the nutrients and oxygen they needed. The cells didn’t die, they just stopped proliferating. This is known as senescence.
5
